This protocol describes the detailed experimental procedure for stem-loop RT-PCR using SYBR Green I. The procedure begins with reverse transcription of total RNA, or small RNA-enriched RNA, miRNA, siRNA, etc. The cDNA is then used as template for real-time PCR with gene specific primers. You may need to modify this protocol if you use different reagents or instruments for real-time PCR.
1.Introduction
The stem-loop RT-PCR method described here is designed to detect and quantify sncRNAs, especially for miRNAs in a fast, specific, accurate and reliable manner. First, a specific stem-loop RT primer is hybridized to the sncRNAs and then reverse transcribed. Next, the RT product is amplified and monitored in real time using a specific forward primer and the universal reverse primer (Fig 1). This method enables sncRNAs expression profiling from as little as 10 pg of total RNA and is suitable for high-throughput sncRNAs expression analysis (Chen et al., 2005; Varkonyi-Gasic et al., 2007; Cheng et al., 2009; Varkonyi-Gasic et al., 2010; Chen et al., 2011; Hurley et al., 2012; Jung et al., 2013; Salone et al., 2015).
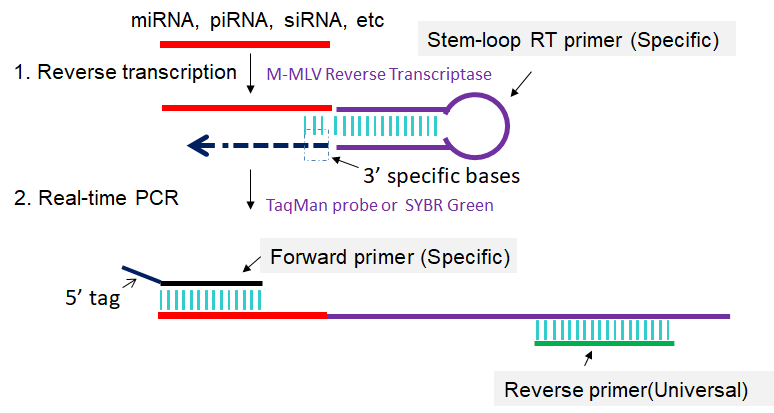
Fig 1. Scheme for stem-loop RT-PCR. 1: Reverse transcription of sncRNA primed by specific stem-loop RT primer. 2: First-strand cDNAs are amplified in PCR by using specific forward primer and reverse primer complementary to stem-loop RT primer.
2. Material and Methods
2.1 Reagents and Equipments
(1) Oligonucleotide Primers. sncRNA specific primers can be designed or retrieved from sRNAPrimerDB (http://www.srnaprimerdb.com.). These primers are ordered from the company (miRNAPrimer: http://www.biootools.net/). All the primers are desalted and both UV absorbance and capillary electrophoresis are used to assess the quality of primer synthesis.
(2) Total RNA, or small RNA-enriched RNA, miRNA, siRNA, etc.
(3) Optical tube and cap strips.
(4) M-MLV Reverse Transcriptase (RNase H-).
(5) SYBR Green PCR master mix.
(6) 50 bp DNA ladder.
(7) Quantitative PCR instrument.
(8) 3% agarose gel.
(9) Agarose gel electrophoresis apparatus.
2.2 Procedure
Reverse Transcription & cDNA Synthesis
Reverse Transcription is carried out with the M-MLV Reverse Transcriptase (RNase H-) for RT-PCR. The following procedure is based on manufacturer’s protocol.
1. Prepare the following RNA/primer mixture in each tube. For each reaction:
| Total RNA | 0.5-1.0 μg |
| RT primer (1 μm earch) | 2 μL |
| DEPC H2O | to 3.5 μL |
2. Incubate the samples at 65 ℃ for 5 min and then on ice for at least 1 min.
3. Prepare reaction master mixture. For each reaction:
| 5 X RT buffer | 4 μL |
| dNTP (10 mmol each) | 4 μL |
| Rnase inhibitor (40 U/μL) | 0.8 μL |
| RTase (200 U/μL) | 1.2 μL |
| DEPC H2O | 6 μL |
4. Add the reaction mixture to the RNA/primer mixture, mix briefly, and then place at room temperature for 2 min.
5. Incubate the tubes in a PCR machine using the following cycling conditions: 16 ℃ for 15 min, 37 ℃ for 20 min, 42 ℃ for 20 min, heat inactivate at 95 ℃ for 5 min, and then chill on ice.
6. Store the 10- or 20- fold dilution 1st strand cDNA at -20 ℃ until use for real-time PCR.
Real-time PCR
The quantitative PCR reaction steps are as follows:
1. Normalize the primer concentrations and mix sncRNA-specific forward and reverse primer pair. Each primer (forward or reverse) concentration in the mixture is 5 pmol/μL.
2. Set up the experiment and the following PCR program on BioRad CFX96 Real-Time PCR. Do not click on the dissociation protocol if you want to check the PCR result by agarose gel. Save a copy of the setup file and delete all PCR cycles (used for later dissociation curve analysis).
| Stage 1: | 94 ℃, 3 min, 1 cycle |
| Stage 2: | 95 ℃, 15 s; 60 ℃ 30 s, 40 cycle |
| Stage 3: | Dissociation analysis |
3. A real-time PCR reaction mixture can be either 50 μL or 25 μL. Prepare the following mixture in each optical tube.
| 25 μL SYBR Green Mix (2X) |
| 0.5 μL cDNA |
| 2 μL primer pair mix (5 pmol/ μL each primer) |
| 22.5 μL H2O |
or
| 12.5 μL SYBR Green Mix (2X) |
| 0.2 μL cDNA |
| 1 μL primer pair mix (5 pmol/μL each primer) |
| 11.3 μL H2O |
4. After PCR is finished, remove the tubes from the machine. The PCR specificity is examined by 3.5% agarose gel using 10 μL from each reaction.
5. Analyze the real-time PCR result. Check to see if there is any bimodal dissociation curve or abnormal amplification plot.
End-point PCR
1. For end-point PCR, it is critical to amplify for an appropriate number of cycles so that 1) the PCR amplification product is readily visible on an agarose gel, and 2) the reaction remains in the exponential phase of amplification. The number of cycles required to meet these two criteria must be empirically determined. As a starting point, we recommend testing 15-25 cycles.
2. Analyze reaction products (10 μL) by electrophoresis on a 3.5% high resolution agarose gel in 1X TAE stained with dye for detecting dsDNA. Gene specific sncRNA amplicons should form discrete ~75 bp bands that are easily distinguished from smaller primer-dimer bands that may be seen in the no-template control reaction.
3. End-point PCR can be used for qualitative determination of differences in the expression of a given sncRNA between two (or more) RNA samples. In addition, a “standard curve” can be constructed to define the approximate magnitude of sncRNA expression differences between samples.
2.3 Primers design
The used example sequences of RNA and DNA oligonucleotides (5’ to 3’) are listed as follows:
Experimental method C: Stem-loop RT-PCR
sRNA ID sRNA Sequence (5'-->3') Length (bp) GC (%)
Let-7a UGAGGUAGUAGGUUGUAUAGUU 22 36.36
RTprimer(specific) GTCGTATCCAGTGCAGGGTCCGAGGTATTCGCACTGGATACGACAACTAT 50 52.00
Fp-1 AACACGCTGAGGTAGTAGGTT 21 47.62 59.34
Fp-2 AACACGTGTGAGGTAGTAGGTT 22 45.45 59.63
Fp-3 AACGCACTGAGGTAGTAGGTT 21 47.62 59.34
Fp-4 AAGAGCGTTGAGGTAGTAGGTT 22 45.45 59.43
Fp-5 AAGCGACCTGAGGTAGTAGGT 21 52.38 60.59
Primer Pair 1
Fp-1 5'-AACACGCTGAGGTAGTAGGTT-3'
|||||||||||||||||||||
5'-AACACGCTGAGGTAGTAGGTTGTATAGTTGTCGTATCCAGTGCGAATACCTCGGACCCTGCACTGGATACGAC-3' (73 bp)
-------**********************
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
|||||||||||||||||||
3'-ACCCTGCACTGGATACGAC-5' Rp
Possible hairpin: 0 kcal/mol (Gibbs free energy)
AACACGCTGAGGTAGTAGGTT
.....................
Possible dimer: -5.5 kcal/mol (Gibbs free energy)
AACACGCTGAGGTAGTAGGTT
|| |||
---TGGGACGTGACCTATGCT
Primer Pair 2
Fp-2 5'-AACACGTGTGAGGTAGTAGGTT-3'
||||||||||||||||||||||
5'-AACACGTGTGAGGTAGTAGGTTGTATAGTTGTCGTATCCAGTGCGAATACCTCGGACCCTGCACTGGATACGAC-3' (74 bp)
--------**********************
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
|||||||||||||||||||
3'-ACCCTGCACTGGATACGAC-5' Rp
Possible hairpin: 0 kcal/mol (Gibbs free energy)
AACACGTGTGAGGTAGTAGGTT
......................
Possible dimer: -3.9 kcal/mol (Gibbs free energy)
-----------AACACGTG
|||
TGGGACGTGACCTATGCTG
Primer Pair 3
Fp-3 5'-AACGCACTGAGGTAGTAGGTT-3'
|||||||||||||||||||||
5'-AACGCACTGAGGTAGTAGGTTGTATAGTTGTCGTATCCAGTGCGAATACCTCGGACCCTGCACTGGATACGAC-3' (73 bp)
-------**********************
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
|||||||||||||||||||
3'-ACCCTGCACTGGATACGAC-5' Rp
Possible hairpin: -0.4 kcal/mol (Gibbs free energy)
AACGCACTGAGGTAGTAGGTT
(((..(((.....)))..)))
Possible dimer: -8.4 kcal/mol (Gibbs free energy)
--AACGCACTGAGGTAGTA
||||||
TGGGACGTGACCTATGCTG
Primer Pair 4
Fp-4 5'-AAGAGCGTTGAGGTAGTAGGTT-3'
||||||||||||||||||||||
5'-AAGAGCGTTGAGGTAGTAGGTTGTATAGTTGTCGTATCCAGTGCGAATACCTCGGACCCTGCACTGGATACGAC-3' (74 bp)
--------**********************
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
|||||||||||||||||||
3'-ACCCTGCACTGGATACGAC-5' Rp
Possible hairpin: 0 kcal/mol (Gibbs free energy)
AAGAGCGTTGAGGTAGTAGGTT
......................
Possible dimer: -3.2 kcal/mol (Gibbs free energy)
----------AAGAGCGTT
|||
TGGGACGTGACCTATGCTG
Primer Pair 5
Fp-5 5'-AAGCGACCTGAGGTAGTAGGT-3'
|||||||||||||||||||||
5'-AAGCGACCTGAGGTAGTAGGTTGTATAGTTGTCGTATCCAGTGCGAATACCTCGGACCCTGCACTGGATACGAC-3' (74 bp)
--------**********************
^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
|||||||||||||||||||
3'-ACCCTGCACTGGATACGAC-5' Rp
Possible hairpin: -1.4 kcal/mol (Gibbs free energy)
AAGCGACCTGAGGTAGTAGGT
.....((((........))))
Possible dimer: -5.4 kcal/mol (Gibbs free energy)
AAGCGACCTGAGGTAGTAGGT
||||
----TGGGACGTGACCTATGC
Total time used: 3.474343759s.
References:
1. Chen C, Ridzon DA, Broomer AJ, Zhou Z, Lee DH, Nguyen JT, Barbisin M, Xu NL, Mahuvakar VR, Andersen MR, Lao KQ, Livak KJ, Guegler KJ. Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res. 2005 Nov 27;33(20):e179.(This paper describes a quantitative stem-loop RT-PCR for the detection of mature miRNAs that is based on TaqMan assays)
2. Varkonyi-Gasic E, Wu R, Wood M, Walton EF, Hellens RP. Protocol: a highly sensitive RT-PCR method for detection and quantification of microRNAs. Plant Methods. 2007, 12;3:12.
3. Cheng A, Li M, Liang Y, Wang Y, Wong L, Chen C, Vlassov AV, Magdaleno S. Stem-loop RT-PCR quantification of siRNAs in vitro and in vivo. Oligonucleotides. 2009, 19(2):203-8. doi: 10.1089/oli.2008.0176.
4. Varkonyi-Gasic E, Hellens RP. qRT-PCR of Small RNAs. Methods Mol Biol. 2010, 631:109-22. doi: 10.1007/978-1-60761-646-7_10.
5. Chen C, Tan R, Wong L, Fekete R, Halsey J. Quantitation of microRNAs by real-time RT-qPCR. Methods Mol Biol. 2011, 687:113-34. doi: 10.1007/978-1-60761-944-4_8. (The experimental principles and procedure of stem-loop reverse transcription (RT)-based TaqMan(®) MicroRNA assays and poly(A) tailing-based and direct RT-based SYBR miRNA assays were introduced in detail.).
6. Hurley J, Roberts D, Bond A, Keys D, Chen C. Stem-loop RT-qPCR for microRNA expression profiling. Methods Mol Biol. 2012, 822:33-52. doi: 10.1007/978-1-61779-427-8_3.
7. Jung U, Jiang X, Kaufmann SH, Patzel V. A universal TaqMan-based RT-PCR protocol for cost-efficient detection of small noncoding RNA. RNA. 2013, 19(12):1864-73. doi: 8.1261/rna.040501.113. (This article describes a universal TaqMan-based probe protocol which can be used to detect any target sequence and demonstrate its applicability for the detection of endogenous as well as artificial eukaryotic and bacterial small RNAs.).
8. Salone V, Rederstorff M. Stem-loop RT-PCR based quantification of small non-coding RNAs. Methods Mol Biol. 2015, 1296:103-8. doi: 10.1007/978-1-4939-2547-6_10.